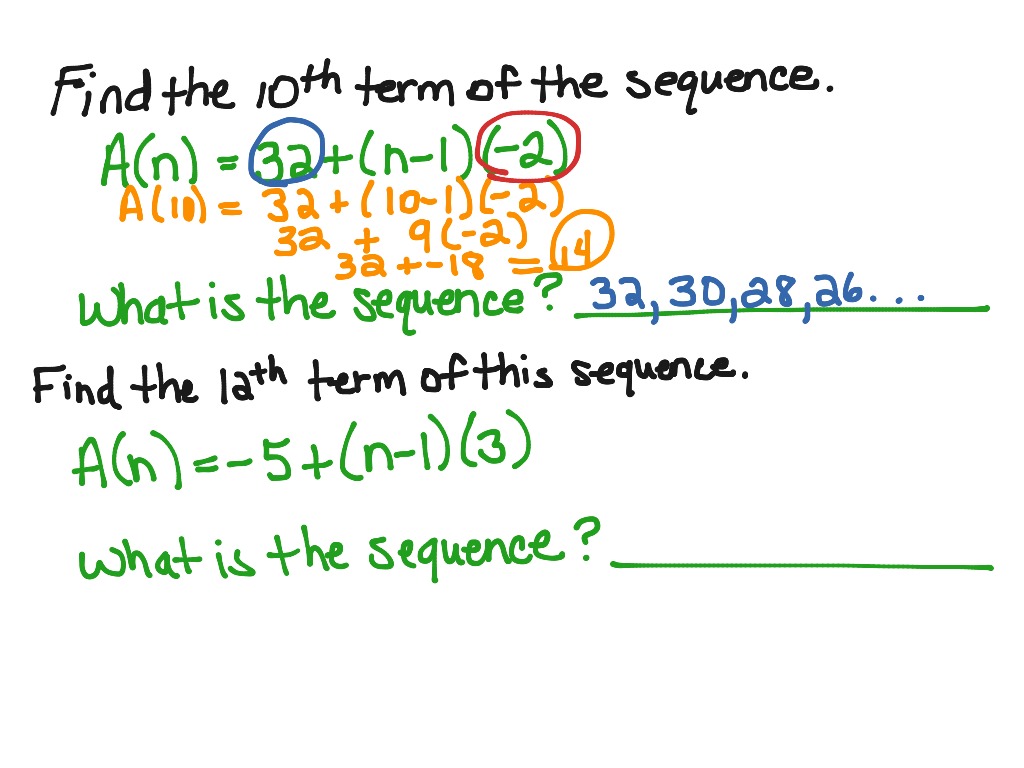
 
Then, proteins are classified in the same family if the remaining HSPs cover at least a given percentage of coverage of the longest protein with a percentage of identity greater or equal to a given threshold (see Figure 1). For each pairwise alignment, the list of High-scoring Segment Pairs (HSPs) is analyzed to exclude HSPs that are not compatible with a global alignment (for details, see ). In practice, protein sequences are compared against each other with BLASTP. Hence, in HOGENOM, proteins are classified in the same family only if they are homologous over their entire length - or almost. The goal of these databases is to allow the study of the evolution of entire proteins considered as a unit, in contrast to databases such as PFAM or PRODOM that aim at studying the domain architecture of proteins. The method presented in this paper was motivated by the development of databases of homologous genes (such as HOGENOM or HOVERGEN ). The choice of these criteria depends on the goal of the clustering. Then, if a pair of sequences (A, B) does not satisfy the criteria, the pair is not considered for the clustering. 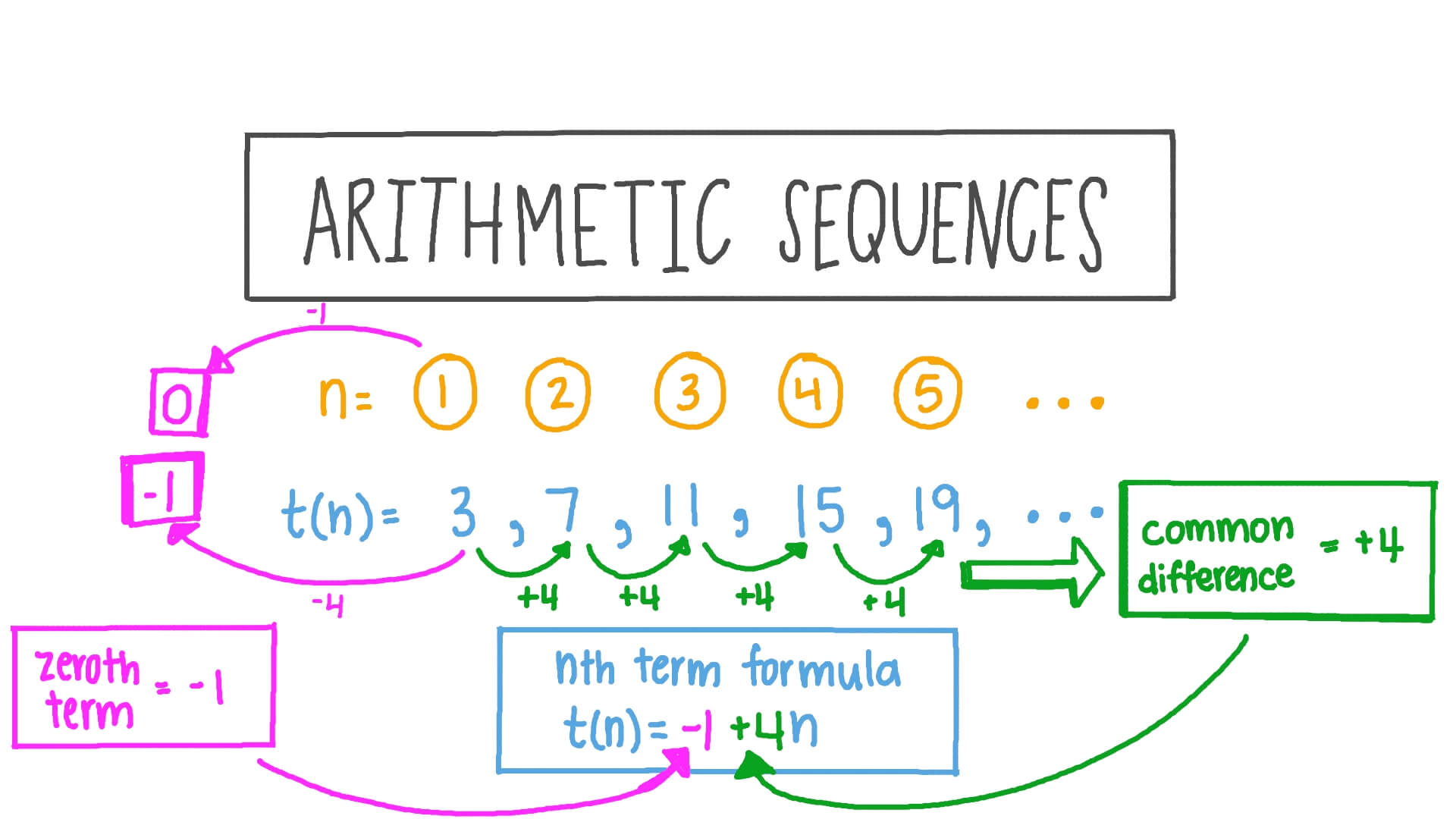
percentage of the length of the sequence that is effectively aligned). Different criteria can be used, separately or in combination (percentage of identity, alignment score or E-value, alignment coverage i.e. 
The choice of the sequence similarity criteria that is used to infer homology is therefore an essential parameter of the single-linkage clustering approach. The principle of the single-linkage clustering is that if sequence A is considered homologous to sequence B, and B homologous to C, then A, B and C are grouped into the same family, whatever the level of similarity between A and C. Modelling Single linkage and filtering with alignment coverage constraints We discuss the interest of SiLiX for the clustering of homologous sequences in huge datasets, possibly in combination with other clustering methods. Moreover, it allows a satisfying quality of clustering. SiLiX outperforms other existing software programs both in terms of speed and memory requirements. Our approach presents several advantages over other clustering algorithms: it is extremely fast, it requires only limited memory and can be run on a parallel architecture - which is essential for ensuring its scalability to large datasets. We evaluated the computational performances and scalability of this method on a very large dataset of more than 3 millions sequences from the HOGENOM phylogenomic database. Finally, we adopt a divide-and-conquer strategy to deal with the quantity of data and design a parallel algorithm whose theoretical complexity is addressed in this paper. This approach enables also an incremental procedure where sequences and similarities are added into the dataset so that it would not be necessary to rebuild the families from scratch. To overcome memory limitations we follow an online framework in which we visit the edges one at a time to update the families dynamically. We model the dataset as a similarity network where sequences are vertices and similarities are edges. In this paper, we present a new approach for the clustering of homologous sequences, based on single transitive links ( single linkage) with alignment coverage constraints and implemented in a software package (called SiLiX for SIngle LInkage Clustering of Sequences). 
With the recent progress of sequencing technologies, there is an urgent need to prepare for the deluge and hence to develop methods able to deal with a huge quantity of sequences. The building of such phylogenomic databases involves three steps that require important computing resources: 1) compare all proteins to each other to detect sequence similarities, 2) cluster homologous sequences into families (that we will call the clustering step) and 3) compute multiple sequence alignments and phylogenetic trees for each family. Thanks to the progress of sequencing projects, this comparative approach can now be applied at the whole genome scale in many different taxa, and several databases have been developed to provide a simple access to collections of multiple sequence alignments and phylogenetic trees. The comparison of homologous sequences and the analysis of their phylogenetic relationships provide very useful information regarding the structure, function and evolution of genes. Proteins can be naturally classified into families of homologous sequences that derive from a common ancestor. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
